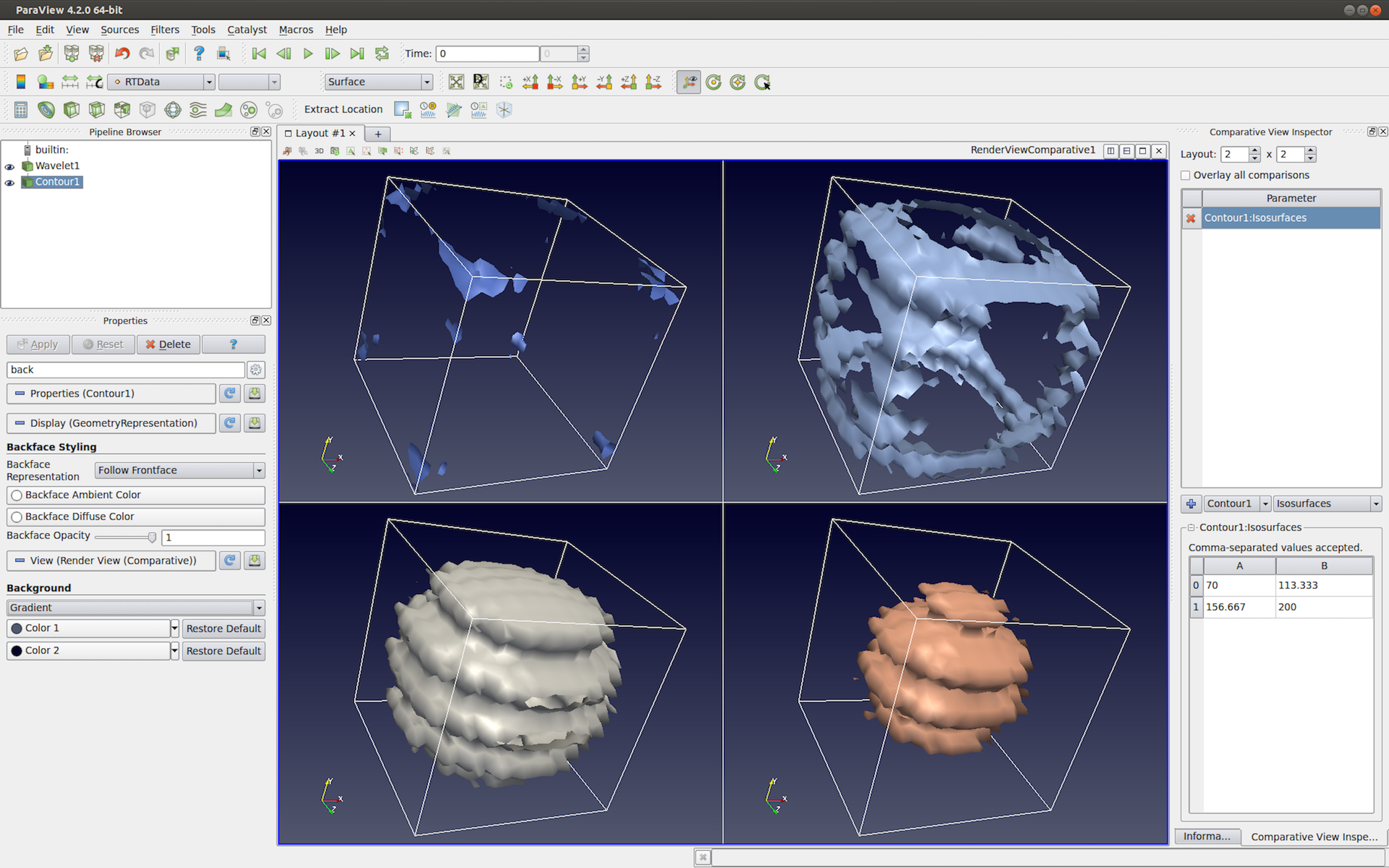 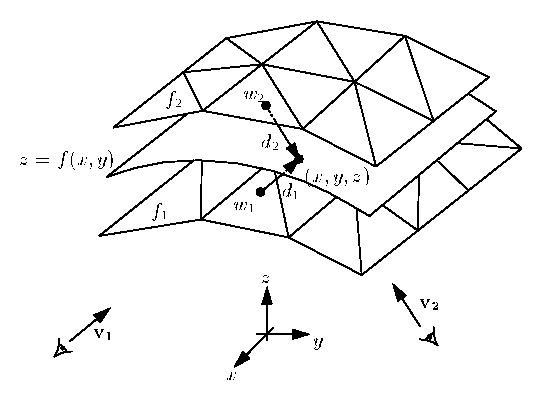
Insight into the properties of each isosurface. This visualization should give you additional In other words, you will visualize the valuesĮach isosurface. Second task is to color map the value of the gradient magnitude on Now that you have identified interesting isosurfacesĬorresponding to major anatomical structures in each dataset, your How did you select the position of clipping planes?.Which isosurfaces look the most interesting? Justify your answer.Include answers to following questions in your report. To facilitate the visual inspection of your results. Identified for each structure and the visual settings associated withĮach visualization. Showing the anatomical structures that you Include in your report high-quality images Initial isovalue to be used by the program, and Optionally import the initial isovalue and initial X, Y, and ZĬlipping plane positions from the command line and initialize the corresponding sliders accordingly.Show the color scale through a color bar.Note: the clipping planes should be applied sequentially: only the half space selected by the X clipping plane will be available to the Y clipping plane, etc. Each plane will defines two half spaces, one of which will not be shown. Of three clipping planes moving along the X, Y, and Z coordinateĪxis, respectively. Provide three additional slider bars GUI to control the position.Ensure consistency between slider bar selection and isosurface.Display next to the slider bar the value currently selected.Provide a slider bar to control the isovalue to be used within.Perform an isosurface visualization of the dataset.Take in input the name of the scalar volume to be visualized.If you do not have it, it can be found on github. Note that this example uses a data file contained in the VTKData distribution. Alternatively, you can refer to the example available here. The implementation of this class shows you how to use vtkScalarBarActor. Note that a helper class named colorbar was included in the PyQt example that I shared with you at the beginning of the semester. This is done with the help of vtkScalarBarActor. You will also add a color bar to your render window to show the meaning of the colors used in your visualization. Which allows you to clip away portions of the VtkClipPolyData filter used with an implicit function called vtkPlanes, Issues, your program will offer interactive control over three clipping planes VtkContourFilter to extract isosurfaces and ties it to a sliderīar GUI to interactively manipulate the isovalue. Yield a set of isovalues that correspond to the majorĪnatomical structures present in each dataset. Scalar dataset using isosurfacing while supporting the interactive modification of theĬorresponding isovalue. The first task consists in visualizing a 3D Finally you will useĬolor mapping to convey quantitative properties of the selected The surfaces to reveal internal structures. The latter will allow you to cut away the occluding part of The surfaces, you will be using transparency and clipping Isovalue, while two additional slider bars will allow you to selectĪ range of gradient magnitudes to act as a filter on theĬonstructed isosurface. Interactively browse the value range in search for the proper Let you select individual values associated with the various Selection of good isovalues, you will create a GUI that will Note that for eachĭataset, the CT and gradient magnitude volumes will offer youĬomplementary means to identify interesting features. 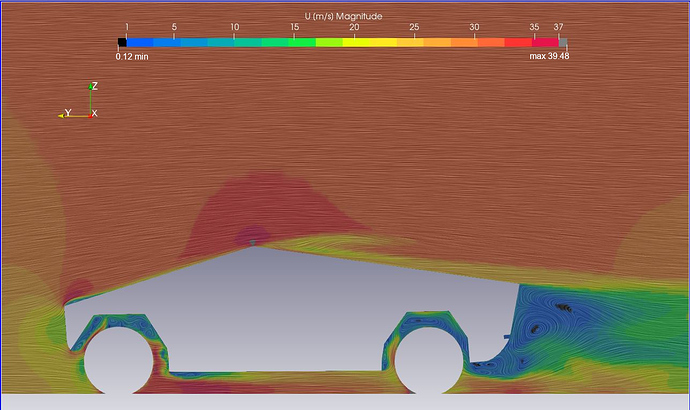
In the footĭataset, you will use isosurfaces to reveal the bone structureĪs well as the skin and surrounding muscles. To identify values that correspond to the skin, skull, and muscles. Your experimentation with the head dataset should allow you Note that other soft tissues ( e.g.,įat or brain) are too difficult to extract. The interesting features in this datasetĬorrespond to skin, bones, and muscles. Your implementation and I expect to see high quality results at full Resolution is mainly provided as a convenience for the testing phase of While the gradient magnitude data is of type float. In each case, the CTĭata is of type unsigned short (16 bits), yielding values between 5, Interested in the magnitude of the derivative. Of a scalar field is the vector-valued derivative of the scalar field. Magnitude of the gradient of the CT image. For each resolution, you willįind a CT volume along with a dataset corresponding to the Halving the size of the original data along each direction (it is Reveal in your visualization the anatomical structuresĪnd low resolution versions are available. Isosurfaces, transparency, clipping planes and color mapping to We will consider a medical imaging dataset corresponding to CT Visualization of three-dimensional scalar fields (3D fields of numbers) using
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
